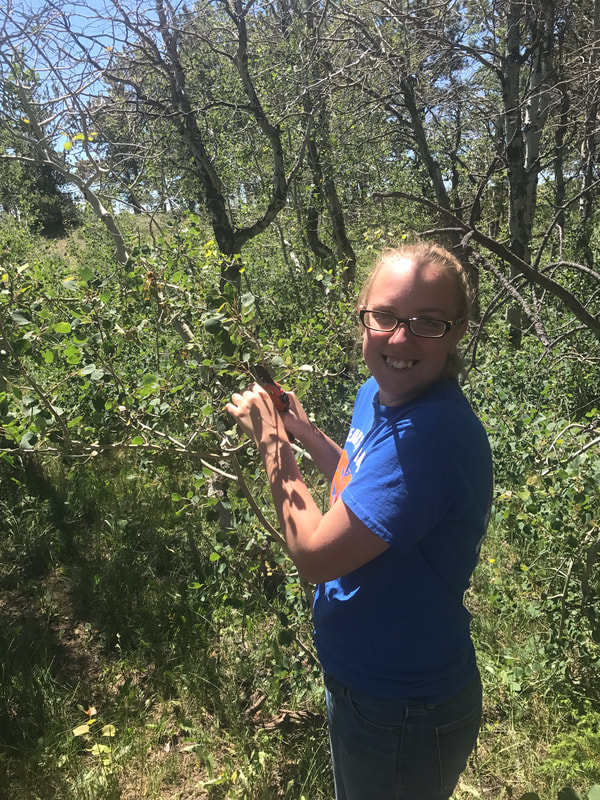
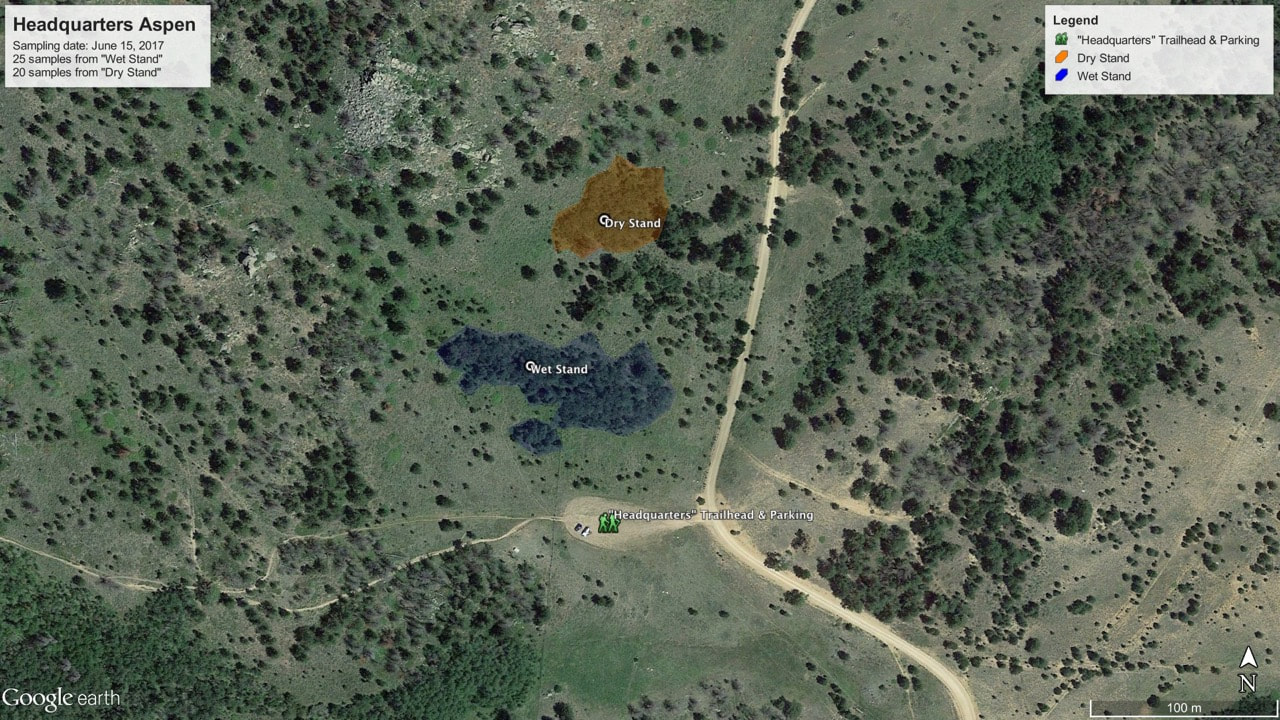
In July of 2017 forty-five P. tremuloides samples were collected at 41.2213, -105.3902 as well as 41.2220, -105.3897. The first coordinates listed were located in a wet stand area. The second coordinates listed were located in a dry stand area. There was a stream located in the wet stand location, whereas the dry stand location did not obtain a stream. This stream was the indication of the different level of moisture between the two stands (Figure 1). Samples were then ground up using liquid nitrogen, mortar, and pestle. DNA was extracted using a modified CTAB protocol, originally by Doyle, J.L., 1987. Agarose gel electrophoresis was then used to observe if DNA was present. Three primers for different loci were optimized. The optimized condition for ORMP-016 was 55.1℃ with two µL of MgCl₂. The optimized condition for ORMP-028 was 55.1℃ with one µL of MgCl₂. The optimized condition for WPMS-020 was 55.1℃ with one µL of MgCl₂. The DNA, which was extracted, was tested with each of the primers. The successful PCR reactions were sorted by individual into a ninety-six well plate (Figure 2), and sent to the University of Arizona for sequencing. Once microsatellite sequence data was received, the data was then analyzed with GENEIOUS software.